Speaker
Description
Introduction
Fluorescence microscopy techniques are widely used in life science research in general and in particular in radiation biology. These techniques involve labelling of various cellular structures with fluorescent dyes followed by observation and image acquisition using fluorescence wide field and laser scanning microscopy. In radiobiological applications, fluorescence microscopy is often employed to detect and enumerate so-called radiation induced foci (RIF). These foci are submicron fluorescent objects in cell culture and tissue section microscopy images that are associated with formation of radiation-induced DNA damage and response to such damage. Common examples of such RIF are $\gamma$H2AX foci that are formed following phosphorylation of H2AX histone at sites of DNA double strand breaks (DSB) or 53BP1 foci that formed after recruitment of 53BP1 protein, which is an important factor of DNA damage repair, at sites of radiation induced DNA damage. For visualisation of these foci, antibodies labelled with fluorescent dyes are used. Microscope image analysis that include detection, enumeration and investigation of RIF kinetics provide valuable information on the extent and repair of radiation-induced DNA damage and widely used in laboratory [1] and clinical [2] research. Given that visual (manual) image analysis is very time consuming and subjected to operator bias, development of software for automated image analysis presents an important step in improving productivity and objectivity of such image analysis. JQuantPro is the latest version of software that initially was developed for quantitative image analysis aiming at automatic detection, counting and analysing properties of RIF in fluorescence microscopy images [3].
Software structure
Apart from performing its major task of detecting and counting RIF, JQuantPro software incorporates a range of additional image and statistical analysis tools that are in part associated with its major task and also can be exploited for solving various general image analysis problems. JQuantPro consists of four modules dedicated for particular stages and types of image analysis. These modules are Image Browser, Object Analysis, Foci Counting and 3D-Counting.
One of the main features of JQuantPro is working with lists of images that allows automatic analysis of multiple images. Image Browser module provide tools for selecting images, browsing through the list of selected images and performing a range of general image manipulation operations such as resampling, normalising, equalising, merging colour planes, building projections, cropping, manipulating image display parameters (contrast, brightness) etc.
The major task of detecting and enumeration of RIF requires implementation of at least two jobs. One of these is the identification in images of cell nuclei that are relatively large objects and termed as “objects” in the context of JQuantPro. Another one is the identification and counting of RIF that are relatively small objects within cell nuclei or larger objects. RIF are termed as “foci” in the context of JQuantPro. The first task is implemented in Object Analysis module that performs identification of objects, associated with cell nuclei, generates object collection for each analysed image in the list and saves object collection files for subsequent foci analysis. Apart from preparing object collection files for foci analysis, Object Analysis module include additional quantitative analysis tools for measuring object properties such as object area, intensity, roundness, building distributions and filtering objects.
The second task is implemented in Foci Counting module that starts from importing object collection files generated at previous step by Object Analysis module and overlaying them on images or colour planes (channels) containing foci signal information. The software allows identification and counting of foci in objects from each object collection file for two additional foci images or foci colour channels, for example $\gamma$H2AX and 53BP1 channels labelled with different fluorescent markers. Apart from counting, a range of foci parameters are measured, as well as frequency distribution of nuclei with respect to foci number, and foci co-localisation analysis is performed if two foci channels are involved. Foci Counting module implements an additional tool for creating groups of images from the list, according to experimental groups in biological experiment, with subsequent statistical analysis generating average parameters for each group.
Image analysis algorithms
JQuantPro has been developed aiming to maximise automation in the process of image analysis. The extent of automation depends on the image quality and complexity, and the uniformity of image properties in the list. It ranges from fully automatic identification of objects to manual drawing and editing of each object with a great level of flexibility allowing the operator to interfere with the automatic analysis process.
Object identification is based on image segmentation at the first step with classical intensity threshold segmentation as a basic algorithm. This algorithm rarely works alone, as for example for low cell density images of cultured cells, and requires some pre-processing of images for better outcome or post processing corrections. For the pre-processing purpose, JQuantPro implements application of noise filters and Top Hat transformation [4, 5]. Top Hat is a classical topological image transformation that efficiently reveals fine structures in 2D image intensity profile and substantially reduce variable background intensity signal.
In addition to intensity threshold segmentation that segments images based on absolute intensity levels, JQuantPro implements H-Dome transformation based segmentation that relies on detection of local intensity maxima [5], thus assisting in identification of objects with different levels of signal intensity.
Objects are defined in JQuantPro as continuous segmented areas of an image. In the majority of cases, post-processing analysis is required that includes filtering of objects based on their size, shape and intensity parameters and splitting of overlapping objects. The software implements an efficient automatic object splitting algorithm based on the analysis of the object size, shape of its boundary and intensity distribution within the object.
Object identification algorithms are dependent on the range of parameters that software attempts to suggest automatically based, for example, on the analysis of image histogram or the comparison of two images with minimal and maximal number of objects. These parameters include the size of Top Hat structuring element, the height of H-Dome transformation, the minimum and maximum size of the object, the minimum roundness of the object etc. The values of these parameters can be fine-tuned by the operator from operator- controlled analysis of one of the images and then can be applied to the automatic analysis of all other images in the list.
Software validation
JquantPro has been validated by comparing results of automatic and operator manual counting of RIF in a few studies [3,6] Figure 1 demonstrates an example of correlation between manual and automatic counting.
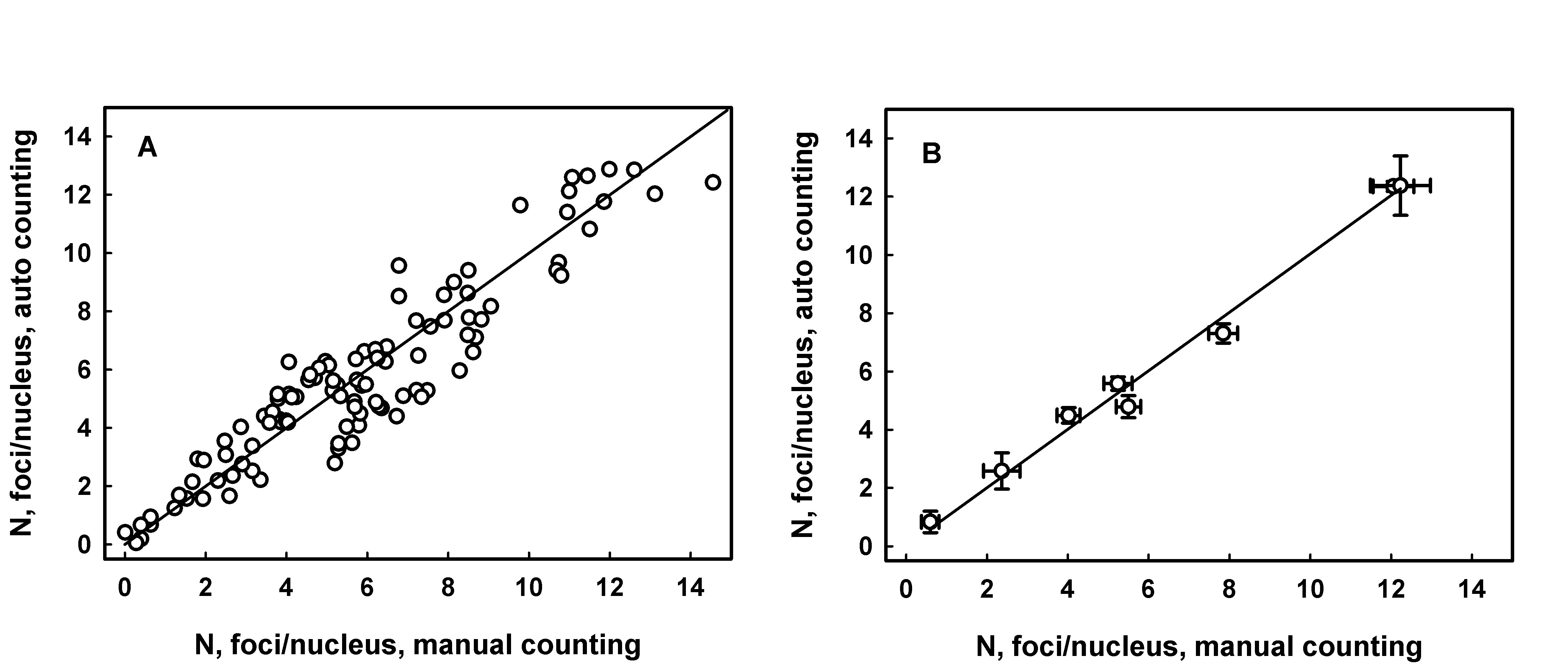
Figure 1. RIF counting analysis in sections of mouse tongue sampled following irradiation of animals with doses from 0.5 to 5 Gy. Panel A: each data point represents average number of foci per nucleus for an individual fluorescence image. The line is generated by linear regression, intercept = 0.26 ± 0.23 (p=0.26, not significant difference from 0), slope = 0.94 ± 0.03, R2 = 0.886. Panel B: each data point represents average number of foci per nucleus for a range of images from an individual experimental group (radiation dose). Error bars indicate the standard error. The line is generated as a result of linear regression analysis, intercept = 0.124 ± 0.308 (p=0.70, not significant difference from 0), slope = 0.99 ± 0.04, R2 = 0.998.
References
[1] P. Lobachevsky, H.B. Forrester, A.Ivashkevich, J. Mason, A.W. Stevenson, C.J. Hall, C.N. Sprung, V.G. Djonov and O.A. Martin. Synchrotron X-Ray Radiation-Induced Bystander Effect: An Impact of the Scattered Radiation, Distance from the Irradiated Site and p53 Cell Status. Frontiers in Oncology, 2021, 11, 685598.
[2] P. Lobachevsky, T. Leong, P. Daly, J, Smith, N. Best, E.R. Thompson, N. Li, I.G. Campbell, R.F. Martin, O.A. Martin. Compromized DNA repair as a basis for identification of cancer radiotherapy patients with extreme radiosensitivity. Cancer Letters, 2016, 383, 212 – 219.
[3] A. N. Ivashkevich, O. A. Martin, A. J. Smith, C. E. Redon, W. M. Bonner, R. F. Martin, P. N. Lobachevsky. γH2AX foci as a measure of DNA damage: a computational approach to automatic analysis. Mutation Research, 711, 49-60, 2011.
[4] M. Van Droogenbroeck, H. Talbot. Fast computation of morphological operations with arbitrary structuring elememnts. Pattern Recognit. Lett., 1996, 17, 1451-1460.
[5] L. Vincent. Morphological grayscale reconstruction in image analysis: applications and efficient algorithms. IEEE Trans. Image Process., 1993, 2 176-201.
[6] L. Jakl, P. Lobachevsky, L. Vokalova, M. Durdik, E. Markova, I. Belyaev. Validation of JCountPro software for efficient assessment of ionizing radiation induced foci in human lymphocytes. International Journal of Radiation Biology, 2016, 92 (12), 766-773.
